Metazoan Phylogeny and Evolution
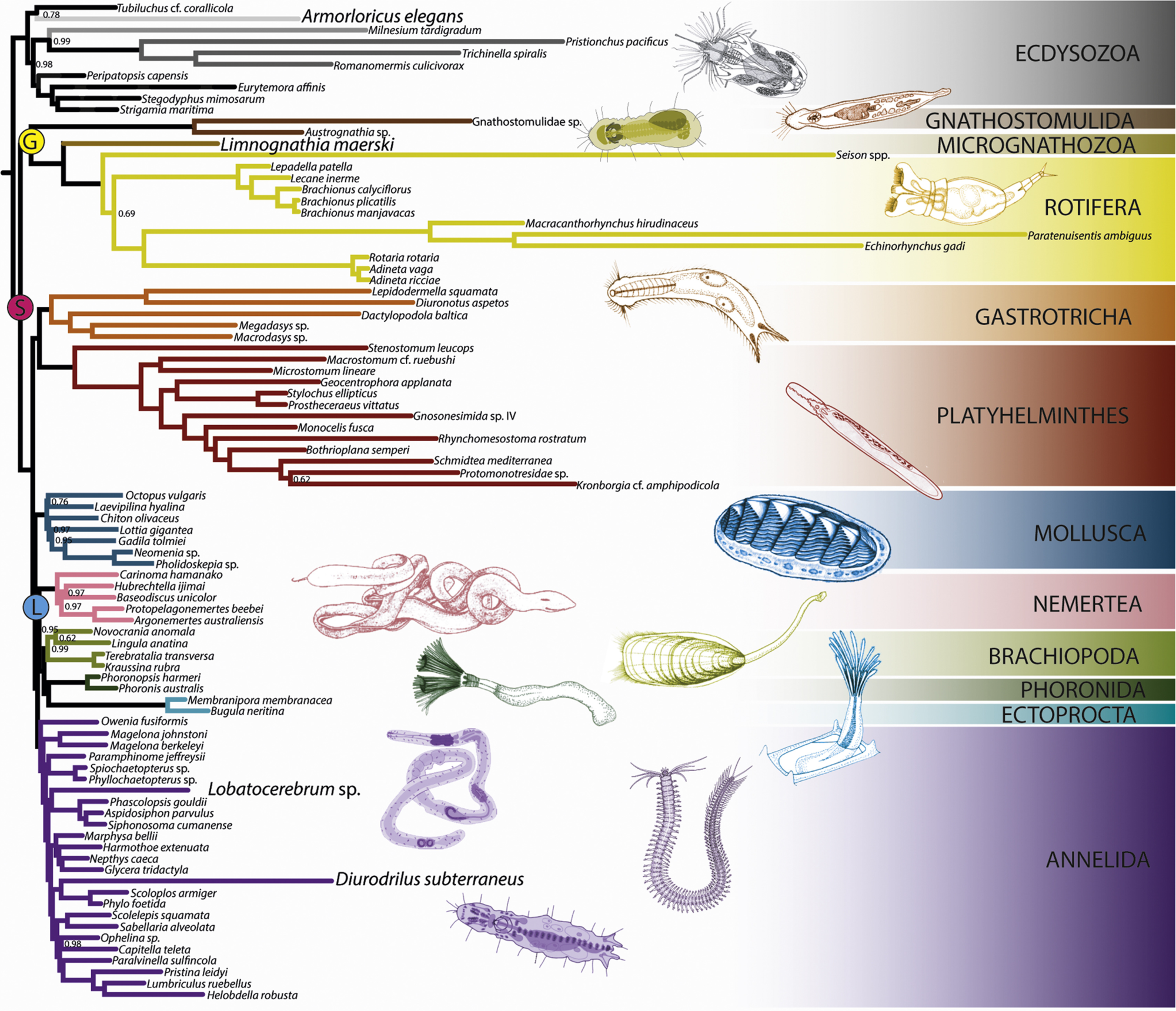
To gain a broad perspective of how animal diversity came to be, we aim at reconstructing the Animal Tree of Life. In collaboration with other research groups, work from the lab has brought up new hypothesis about animal evolution, such as the position of Ctenophora as the sister group to all other metazoans (Dunn et al. 2008), and the position of the elusive placozoans as the sister group to cnidarians (Laumer et al. 2018). We apply new large scale data to study metazoan relationships and to the sampling of small, but key lineages, targeting the root of bilaterian animals (Hejnol et al. 2009) and relationships among the diverse spiralian phyla (Laumer et al. 2015).
Phylogenomics of Invertebrate Clades
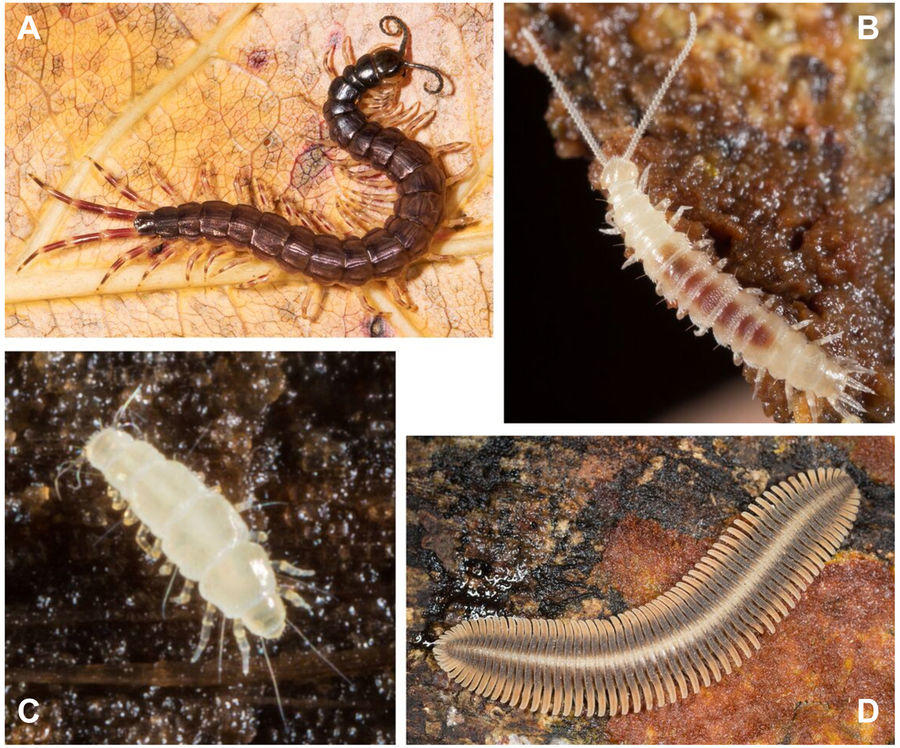

A major goal in the lab is to elucidate the evolutionary relationships of large metazoan lineages. A lot of this work is focused on two of the most diverse animal clades, arachnids and mollusks (more on sections below), but we also work on a diversity of other groups, such as onychophorans, annelids, millipedes and centipedes, nemerteans, crustaceans, platyhelminthes, tardigrades and more. We use RNAseq to produce large phylogenomic datasets of hundreds to thousands of genes to resolve deep divergences in the animal tree. For example, Schwentner et al. (2017) investigate the origins of insects, which are nested within crustaceans, to determine which crustacean lineage is the sister-group to insects and springtails.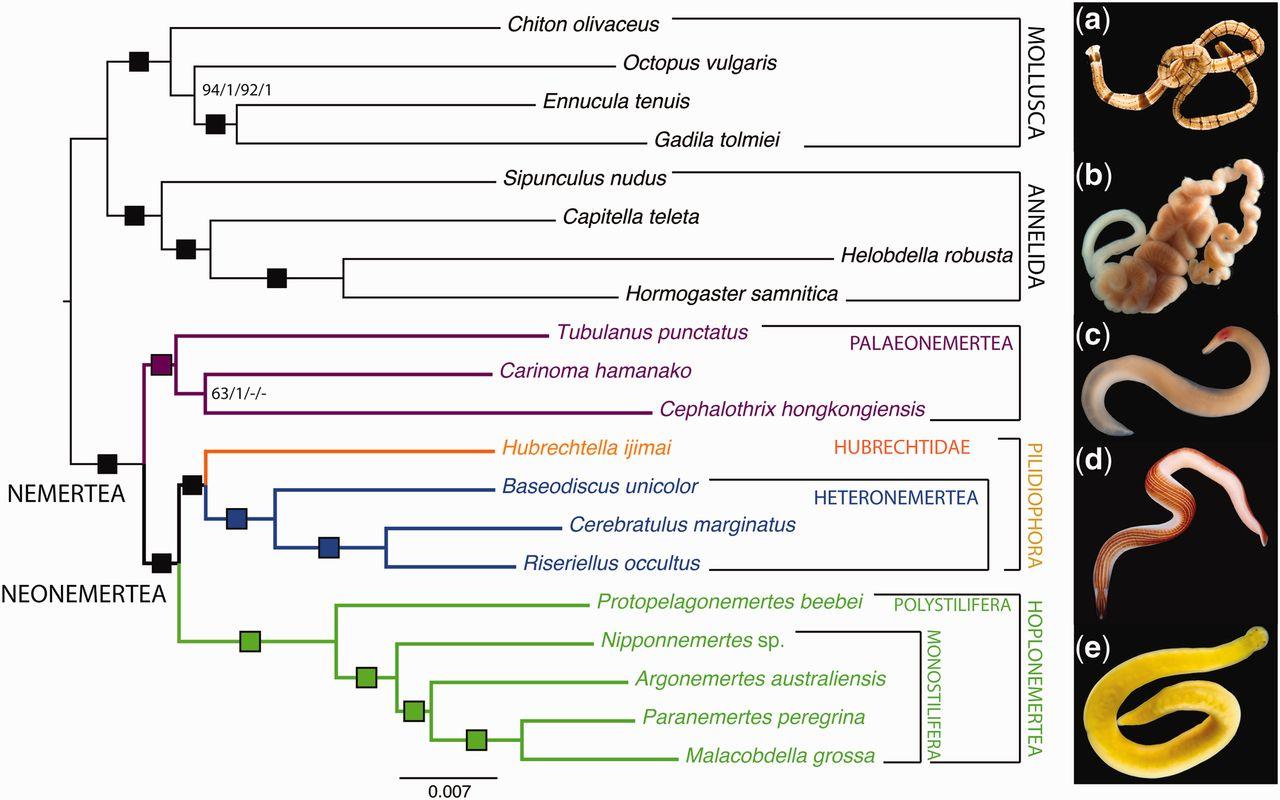

We also incorporate fossils (with morphology data or ages only) to estimate the time of divergence between taxa and discuss events like the terrestrialisation of arthropod lineages, as done for myriapods in Fernandez et al. (2018). Further phylogenomic work has incorporated underrepresented lineages to clarify relationships among annelids (Andrade et al. 2015), nemerteans (Andrade et al. 2014), and to bring novel insights about the evolutionary transition between free-living and vertebrate-parasitic lifestyles in flatworms (Laumer et al. 2015). We also use a variety of analytical methods to explore systematic issues like matrix occupancy and data heterogeneity in phylogenomic datasets.
Biogeography and Phylogeography of Terrestrial and Marine Invertebrates
Phylogenenies are useful tools in evolutionary biology, and one of the major ways in which we use them in the Giribet Lab is to explore biogeographic and phylogeographic patterns in invertebrates. For our work on historical biogeography, we generate time-calibrated phylogenies using known fossils and molecular dating methods. We then compare the 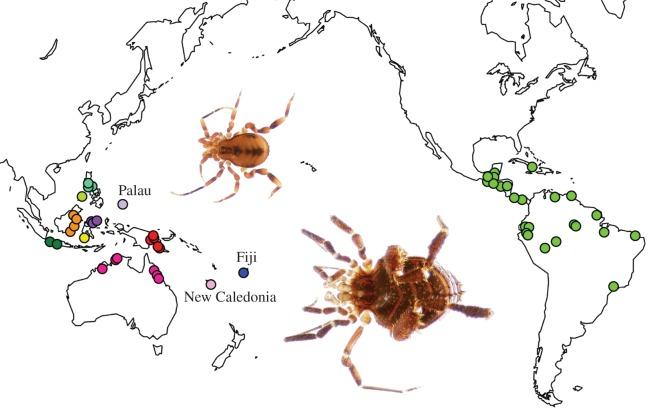 timing and order of when clades diverged to geologic and climatic events in the history of Earth to try and understand why different species live where they do, and why some places on earth are more diverse than others. Part of this research has focused on soil invertebrates with limited dispersal capacity, such as harvestmen (e.g. Giribet et al. 2016, Sharma and Giribet 2012), velvet worms (e.g. Murienne et al. 2014), and centipedes (e.g. Vélez et al. 2011). On the marine side, we have targeted oceanic routes of dispersal of gastropods across hundreds of millions of years (Cunha et al. 2019), as well as the role of abiotic factors (e.g. ocean currents, temperature) and life history strategies on phylogeographical patterns, for example in flatworms (Riesgo et al. 2017) and molluscs (Fernández et al. 2014).
timing and order of when clades diverged to geologic and climatic events in the history of Earth to try and understand why different species live where they do, and why some places on earth are more diverse than others. Part of this research has focused on soil invertebrates with limited dispersal capacity, such as harvestmen (e.g. Giribet et al. 2016, Sharma and Giribet 2012), velvet worms (e.g. Murienne et al. 2014), and centipedes (e.g. Vélez et al. 2011). On the marine side, we have targeted oceanic routes of dispersal of gastropods across hundreds of millions of years (Cunha et al. 2019), as well as the role of abiotic factors (e.g. ocean currents, temperature) and life history strategies on phylogeographical patterns, for example in flatworms (Riesgo et al. 2017) and molluscs (Fernández et al. 2014).
 timing and order of when clades diverged to geologic and climatic events in the history of Earth to try and understand why different species live where they do, and why some places on earth are more diverse than others. Part of this research has focused on soil invertebrates with limited dispersal capacity, such as harvestmen (e.g. Giribet et al. 2016, Sharma and Giribet 2012), velvet worms (e.g. Murienne et al. 2014), and centipedes (e.g. Vélez et al. 2011). On the marine side, we have targeted oceanic routes of dispersal of gastropods across hundreds of millions of years (Cunha et al. 2019), as well as the role of abiotic factors (e.g. ocean currents, temperature) and life history strategies on phylogeographical patterns, for example in flatworms (Riesgo et al. 2017) and molluscs (Fernández et al. 2014).
timing and order of when clades diverged to geologic and climatic events in the history of Earth to try and understand why different species live where they do, and why some places on earth are more diverse than others. Part of this research has focused on soil invertebrates with limited dispersal capacity, such as harvestmen (e.g. Giribet et al. 2016, Sharma and Giribet 2012), velvet worms (e.g. Murienne et al. 2014), and centipedes (e.g. Vélez et al. 2011). On the marine side, we have targeted oceanic routes of dispersal of gastropods across hundreds of millions of years (Cunha et al. 2019), as well as the role of abiotic factors (e.g. ocean currents, temperature) and life history strategies on phylogeographical patterns, for example in flatworms (Riesgo et al. 2017) and molluscs (Fernández et al. 2014).
Biodiversity Discovery
In the Giribet Lab we are interested in documenting and understanding the life on Earth. Participating in field expeditions to different parts of the world (see below) in order to discover, describe and place new species in a phylogenetic context is one of the main topics that drive our research. In addition to traditional techniques (such as light photography and Scanning Electron Microscopy, SEM) used to study the morphology of our favorite organisms, we are interested in developing novel methods. New techniques that we are currently using in the lab include confocal microscopy and micro computed tomography.
Research on Mollusks
While half of the Giribet lab is investigating the 8-legged life on land, the other half is investigating the one-footed, or multi armed life under the waves. Systematics, population genetics and species delimitation of mollusks is a major interest and theme of ongoing research, primarily in bivalves and gastropods. We are additionally interested in questions of deeper mollusk evolution, such as the relationships of the major extant classes.
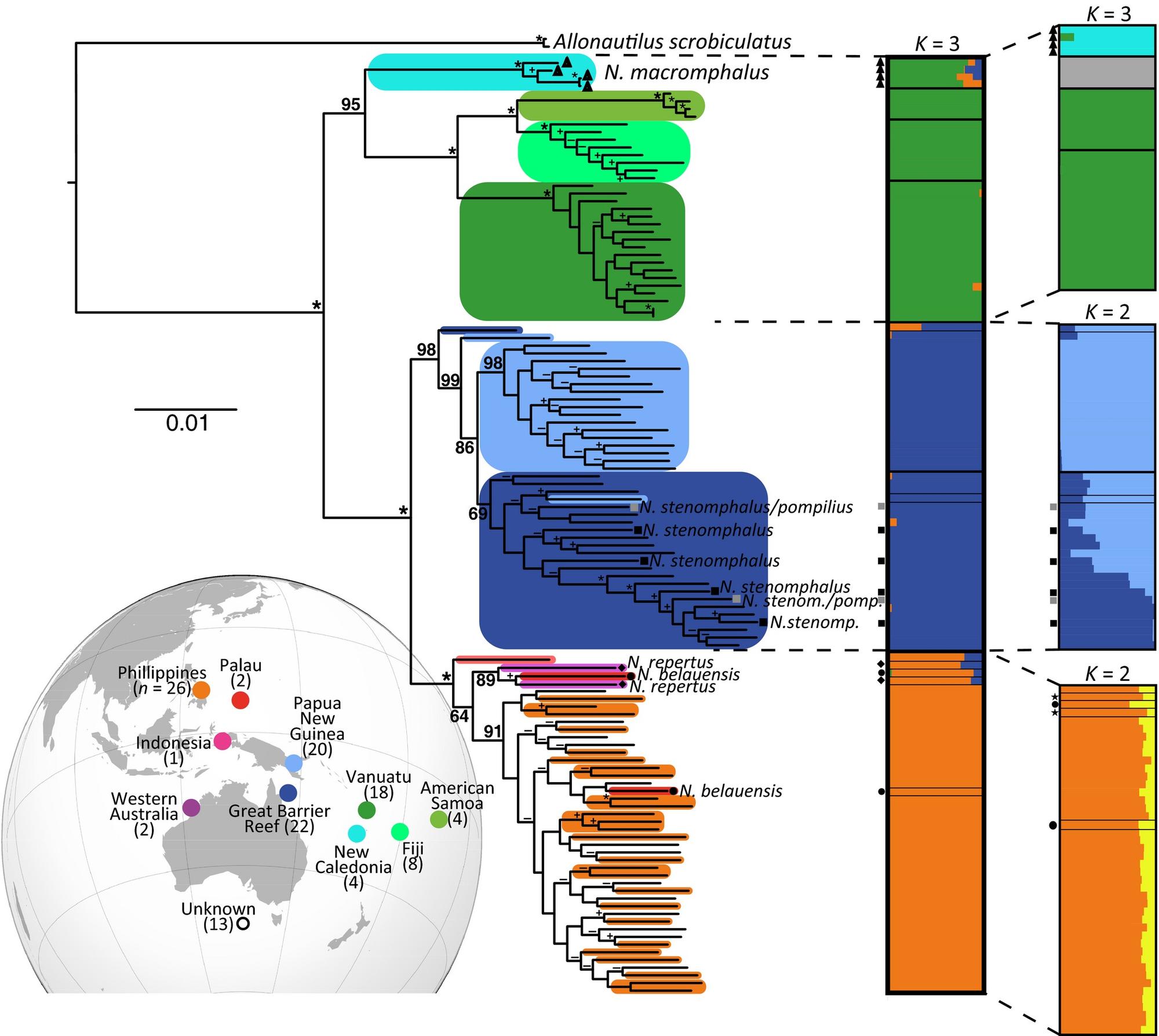
We use diverse techniques, such as ddRad-Seq to investigate the taxonomy, population genetics and effective population sizes of Nautilus (Combosch et al. 2017), and RNAseq to elucidate the relationships within and among major clades of bivalves (Lemer et al. 2019) and gastropods (Cunha & Giribet 2019), finally resolving long-standing questions of phylogeny.
Research on Arachnids
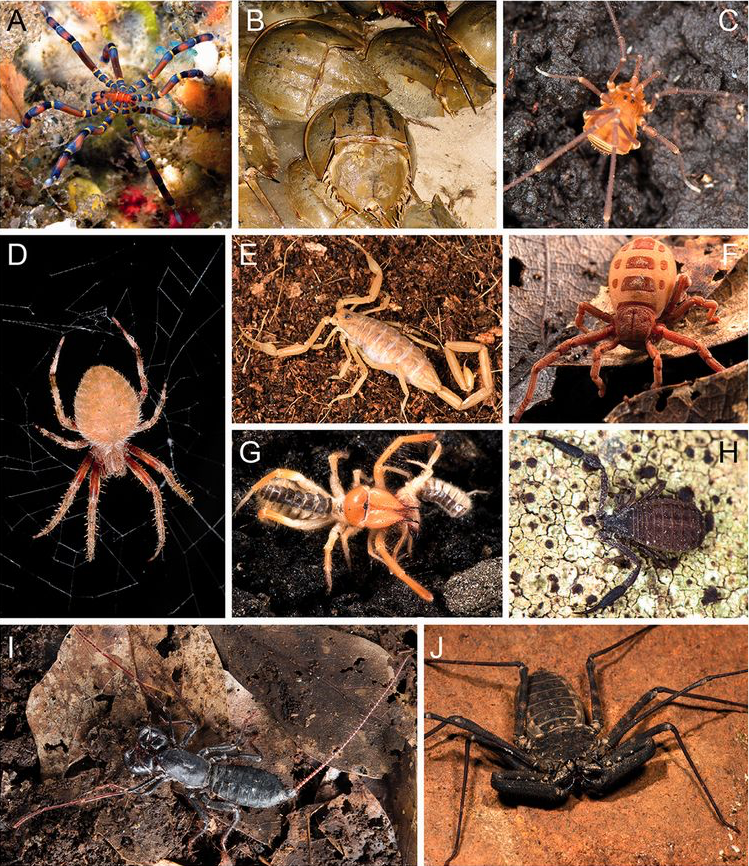

The overall aim of our arachnid research is to reconstruct phylogenies which are then utilized to explore questions relating to phylogenetics, taxonomy, and evolutionary biology. We are interested in arachnid systematics at multiple taxonomic levels using molecular techniques to reconstruct relationships among arachnid orders (Sharma et al. 2014), within various orders e.g. spiders (Fernández et al. 2018), Opiliones (Fernández et al. 2017), Scorpiones (Sharma et al. 2015), all the way down to species level analyses that include new species discovery and descriptions (Giribet et al. 2017). Most of these studies utilize transcriptomes to reconstruct phylogenomic relationships among lineages, however, we are also incorporating ultraconserved elements in our phylogenomic approaches. One major focus of our arachnid research has been on the order Opiliones (harvestmen). This research has been worldwide in geographic scope and spans all of harvestmen diversity, with recent attention now focusing on the early-diverging Gondwanan Laniatores family Triaenonychidae, particularly the New Zealand fauna.
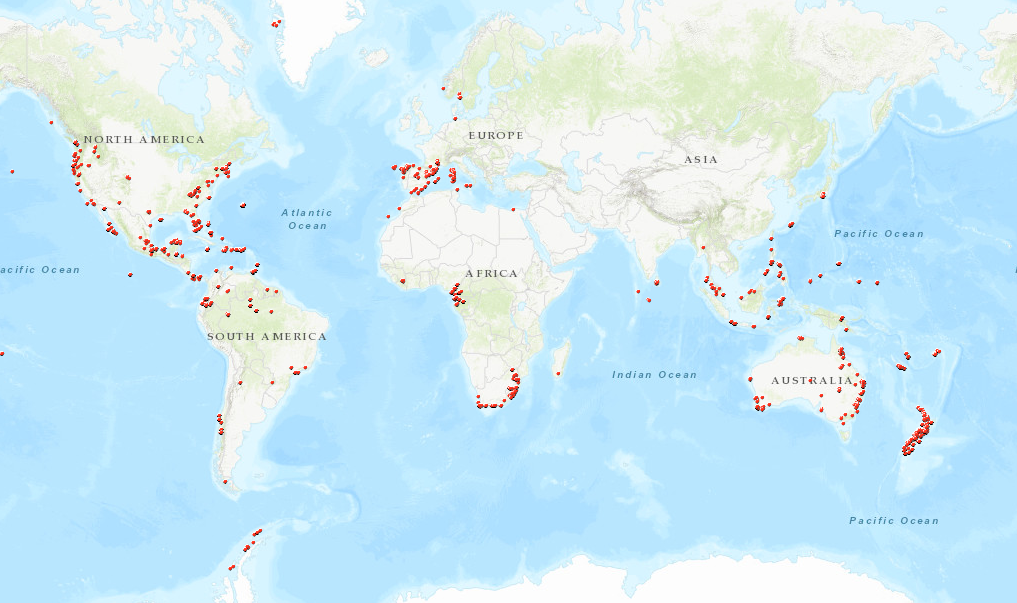
Field work localities through the years by members of the Giribet Lab
Fieldwork is an important component of biodiversity research. We strive to sample organisms from the most diverse places in the world, going wherever the animals are. Another important part of this work is processing the specimens and depositing them in natural history collections, so that information is broadly accessible and linked to other databases. We do this in the Museum of Comparative Zoology, and you can search the MCZ online database to find collecting information, photos, genetic data and much more for our specimens!